Genetic recombination

Genetic recombination is the name given to a group of reactions during which cellular machinery uses DNA to alter or "recombine" with a similar (homologous) sequence. The process involves pairing between complementary strands of DNA, and results in a physical exchange of chromosome material. Genetic information is recombined by the cell for several reasons including the repair of damaged DNA, and the production of population variability during sexual reproduction. In some cases, recombination is known to change genes, adding new alleles to the population.
Creationists generally believe that this mechanism was designed to generate the tremendous variety that is evident within each Biblical kind, whereas evolutionists attribute such variability ultimately to random mutagenesis.[1] However, many creationists contend that recombination processes add nothing new to the gene pool. Jonathan Sarfati states:
| “ | Biologists have discovered a whole range of mechanisms that can cause radical changes in the amount of DNA possessed by an organism. Gene duplication, polyploidy, insertions, etc., do not help explain evolution, however. They represent an increase in amount of DNA, but not an increase in the amount of functional genetic information—these mechanisms create nothing new.[2] | ” |
Background
The position at which a gene is located on a chromosome is called a locus. In a given individual, one might find two different versions of this gene at a particular locus. These alternate gene forms are called alleles. During Meiosis I, when the chromosomes line up along the metaphase plate, the two strands of a chromosome pair may physically cross over one another, and during these events genetic recombination if performed by the cell.[3]
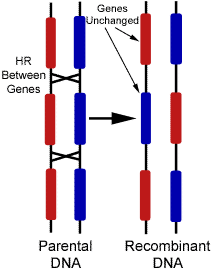
Recombination results in a new arrangement of maternal and paternal alleles on the same chromosome. Although the same genes appear in the same order, the alleles are different. This process explains why offspring from the same parents can look so different. In this way, it is theoretically possible to have any combination of parental alleles in an offspring, and the fact that two alleles appear together in one offspring does not have any influence on the statistical probability that another offspring will have the same combination. This theory of "independent assortment" of alleles is fundamental to genetic inheritance. However, having said that, there is an exception that requires further discussion.[3]
The frequency of recombination is actually not the same for all gene combinations. This is because recombination is greatly influenced by the proximity of one gene to another. If two genes are located close together on a chromosome, the likelihood that a recombination event will separate these two genes is less than if they were farther apart. Linkage describes the tendency of genes to be inherited together as a result of their location on the same chromosome. Linkage disequilibrium describes a situation in which some combinations of genes or genetic markers occur more or less frequently in a population than would be expected from their distances apart. Scientists apply this concept when searching for a gene that may cause a particular disease. They do this by comparing the occurrence of a specific DNA sequence with the appearance of a disease. When they find a high correlation between the two, they know they are getting closer to finding the appropriate gene sequence.[3]
Evolutionary Assumptions
The theory of evolution has led to the assumption that recombination originally occurred by mistake, instead of being an intelligently designed process. During sexual reproduction, gametes (egg, sperm) are produced during a cell division process called meiosis. Prior to meiotic division, homologous chromosomes unite at the cell axis before dividing to opposite poles. It is believe that this homologous pairing was originally performed simply to insure an equivalent division of genetic information. But, an exchange of DNA accidentally occurred during this process, which provided beneficial variability and was naturally selected to became a regular part of gamete formation. It remains generally assumed that recombination events are rather random, and therefore, the phenotypes produced by these reaction are also.[1]
The DNA used for meiotic recombination possess homology or sequences that are very similar, and also code for variations of the same characteristic. Before the chromosomal DNA is distributed into new daughter cells, the homologues pair and are spliced together at multiple locations. During these interactions, entire regions and many genes are frequently exchanged. These genetic crossovers are commonly used to deduce the relative position of genes on chromosomes, and thereby construct genetic maps. Contrary to evolutionary assumptions, the purpose for these manipulations has been preprogrammed by the creator. Offspring are always genetically unique due to recombination, but we have only been able to recognize the most obvious products of these reactions, and the desired outcomes remain largely theoretic. However, it is now clear that recombination is a powerful source of new alleles[1]
Our knowledge of recombination comes predominantly from the bacteria E. coli, and its effect during sexual reproduction (meiosis) has been studied mostly using lower eukaryotes such as baker's yeast, as well as fruit flies. Recent work with mice has provided additional information from mammals, and shown that substantial differences exist between unicellular and multicellular organisms. However, as with most cellular housekeeping mechanisms, the basic details and many genes involved in homologous recombination (HR) appear conserved among the multitude of life forms on earth.[4] It is now widely recognized that genetic editions through HR are part of a highly coordinated process involving a cascade of specific macromolecule interactions,[5] and controlled by highly organized regulatory systems.[6] In particular, the induction of recombination during meiosis is reliant upon several genes, and is regulated by a complex network of cell signaling mechanisms.[7]
Non-Random Recombination
Since their discovery and use in the construction of genetic maps, it was assumed that gene crossovers during meiosis occurred at random intervals along chromosomes. It was believed that the frequency of gene crossovers was directly related to the distance between genes, but a variety of discoveries have illustrated the existence of differential recombination rates and patterns, and forced a revision of map distances. It is now a well-known fact that recombination frequency is not constant in any one particular cell. Reactions occur more frequently in some regions of the genome than in others with variations of several orders of magnitude observed. These hyperactive regions have been termed as "hot spots" as opposed to inert "cold spots" where little to no exchange is found.[8]
The frequencies of recombination events are also nonrandom. The rates are found to be significantly higher when comparing germ-line with somatic cell types. For example, mitotic recombination frequencies in the fungus Ustilago maydis have been estimated at 2.9 x 107; whereas, in meiosis the rates are closer to 1.9 x 103. Sex-specific differences in recombination frequency have also been elucidated. Standard linkage analysis was used to confirm that females have a higher recombination rate than males, and males recombine preferentially in the distal regions of the chromosome. These and other techniques were also separately used to establish the existence of significant inter-individual variation in recombination over short intervals.[9] Still other researchers have demonstrated background effects on the frequency of recombination using immunostaining techniques to assess meiotic exchange patterns. It has now been found in many cases, that crossover events are non-randomly distributed and display positive interference.[10]
In addition to exchanges during cell division, HR is involved with many other forms of genomic DNA editing. For example, recombination is induced or shut off as a preprogrammed cell function during differentiation and development. It is also used to perform error-free DNA repair, which in this case serves to prevent unintentional variability. In fact, HR maintains the integrity of the genome through the correction of several different types of DNA damage.[7] Homologous recombination is stimulated by double-stranded breaks during any stage of the cell cycle, and is also responsible for performing deletions, duplications, and translocations between dispersed homologous, which are frequently a response to stress.[11] The specific details or exact sequence homology required for recombination remain largely unknown, but the plethora of functions accomplished by these reactions has elevated them to the position of master mechanic responsible for virtually all forms of sequence editing and maintenance.
New Alleles
There is an interesting new class of HR only recently recognized that shares common mechanisms with meiotic crossovers, and is likely responsible for the formation of new alleles. The process known as gene conversion uses template DNA to edit active sequences. During this process, pseudo genes previously referred to as junk DNA is frequently used to make these changes.[12] Gene conversion can be easily distinguished from crossovers in most cases because only one of the homologues are altered. It has now been thoroughly documented that mitotic recombination via gene conversion is able to create genetically altered cells, and researchers have suggested that this process can generate a gene with novel functions by rearranging various parts of the parental reading frames.[13] DNA is also repaired through conversion when an intact copy from the sister chromatid or homologous chromosome is used to replace the damaged region. Gene conversion is now understood to be responsible for performing many alterations that were previously attributed to mutations or other repair mechanisms.

Variable Genes
Diversification within a population occurs because the genes involved with the production of characteristics exist as a variety of alleles, and therefore traits are polymorphic or available in more than one form. Closely related species are commonly found with extremely high numbers of alleles. For example, the cystathionine ß-synthase gene locus has been intensely studied in humans, and Exon 8, in particular, has a high frequency of single nucleotide alterations. It is estimated that approximately 5% of human Caucasians possess variations in this region.[14] Evolutionists generally assume that new alleles are the result of random mutations that have accumulated gradually over millions of years. However, living populations have been tested only decades following severe genetic bottlenecks to find surprisingly high genetic diversity. This strongly suggests a mechanism for rapidly restoring variability, and the yet this possibility has not yet been adequately explored.18 However, an explanation for this continued diversity was suggested when it was discovered that many genes in every genome are highly diverse (hypervariable) in comparison to others.
Not all genes are variable. The majority of genes in the genome are involved with housekeeping functions, and are commonly found unchanged even when comparing vastly different organisms. In contrast, variable genes change significantly from one generation to the next and show nonrandom patterns within any given gene.[15] The characterization of variable genes to date suggests overwhelmingly that this diversity is systematically produced through gene conversion while under tight cellular control. For example, variable genes have hot and cold spots of activity similar to those found among gene crossovers in meiosis.[16] They also frequently have greater diversity than the neutral regions between reading frames.[17] It has likewise become evident that variable genes retain codons at specific locations within the variable region. [18] A preponderance of non-synonymous substitutions over synonymous has provided even further evidence against randomness.[19] It is becoming increasingly questionable that variability is the result of random mutations as commonly claimed by evolutionists.
Adaptation
Adaptation to a particular habitat or niche involves largely uncharacterized modifications of the genome, and much of what we've learned about genetic heredity has come from theorists who do not believe the cell was designed to perform such changes with intent. The ability of the cell to produce new alleles has probably remained misunderstood for so long because the products of these reactions are being attributed to a source that is independent of cellular purpose (mutations). The mechanisms behind this type of gene conversion are not yet understood, but clearly illustrate the ability of the cell to specifically edit genes, and thereby rapidly multiply the number of alleles in a population. Further characterization should prove to be valuable evidence that cellular design governs the production of genetic variability, and adaptive evolution that occurs as a result.
References
- ↑ 1.0 1.1 1.2 .Genetic Variability by Design by Chris Ashcraft. Journal of Creation 18(2) 2004.
- ↑ Sarfati, Jonathan. Refuting Evolution 2 Chapter 5 - Argument: Some mutations are beneficial. Greenforest AR: Master Books, 2002. (p104)
- ↑ 3.0 3.1 3.2 What is a Cell? by the National Center for Biotechnology Information
- ↑ Regulation of meiotic recombination and prophase I progression in mammals Cohen P.E. & Pollard J.W. BioEssays 23:996-1009 (2001)
- ↑ Cascades of Non-covalent Protein-protein and Protein-DNA Interactions for Homologous DNA Recombination Takehiko Shibata. RIKEN Review 46:24-28 (2002)
- ↑ Hierarchic Regulation of Recombination Kunihiro Ohta. RIKEN Review 41:28-29 (2001)
- ↑ 7.0 7.1 Homologous genetic recombination as an intrinsic dynamic property of a DNA structure induced by RecA/Rad51-family proteins: a possible advantage of DNA over RNA as genomic material Shibata, T., Nishinaka, et al. Proc. Natl. Acad. Sci. U.S.A. 98(15):8425-8432 (2001)
- ↑ Meiotic recombination hotspots Lichten, M. & Goldman, A.S.H. Annu. Rev. Genet. 29:423-444 (1995)
- ↑ Counting Cross-overs; Characterizing Meiotic Recombination in Mammals Terry Hassold. Human Molecular Genetics 9(16):2409-2419 (2000)
- ↑ Genetic control of Mammalian meiotic recombination. I. Variation in exchange frequencies among males from inbred mouse strains Koehler KE, Cherry JP, Lynn A, Hunt PA, Hassold TJ. Genetics 162(1):297-306 (2002)
- ↑ Homologous Recombination as a Mechanism for Genome Rearrangements: Environmental and Genetic Effects Alexander Bishop. Human Molecular Genetics 9(16):2427-2434 (2000)
- ↑ The chicken B cell compartment Weill JC, Reynaud CA. Science 238(4830):1094-1098 (1987)
- ↑ Functions of Homologous DNA Recombination Takehiko Shibata. RIKEN Review 41:21-23 (2001)
- ↑ Allozyme evidence for crane systematics and polymorphisms within populations of sandhill, sarus, Siberian and whooping cranes. Dessauer, H. C., G. F. Gee, and J. S. Rogers. Molecular Phylogenetics and Evolution 1:279-288 (1992)
- ↑ Creation of immunoglobulin diversity by intrachromosomal gene conversion. Thompson, C. B. Trends in Genetics 8:416-422 (1992)
- ↑ The targeting of somatic hypermutation Jolly, C.J. et al. Seminars in Immunology 8:159-168 (1996)
- ↑ Gene conversion generates hypervariability at the variable regions of kallikreins and their inhibitors. Ohta, T. and C. J. Basten. Molecular Phylogenetics and Evolution 1:87-90 (1992)
- ↑ Position-specific codon conservation in hypervariable gene families Conticello, S. G., Y. Pilpel, G. Glusman, and M. Fainzilber.Trends Genet. 16:5759 (2000)
- ↑ Mechanisms for Evolving Hypervariability: The Case of Conopeptides Conticello, S. G., Gilad, Y., Avidan, N., Ben-Asher, E., Levy, Z., Fainzilber, M. Mol Biol Evol 18:120-131 (2001)
| |||||||||||||||||