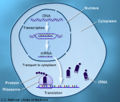
Gene expression
Gene expression is the process by which the information encoded in a gene is converted into protein or some form of RNA. The process has two steps called transcription and translation. First the DNA sequence transcribed into RNA and then usually, but not always, translated into protein.
A gene is said to be expressed when its information is transcribed onto a messenger RNA with instructions for a specific protein to be made. The messenger RNA is essentially a copy of the gene that is delivered to a cellular machine (ribosome), which then uses the information to assemble a specific sequence of amino acids. This is known as the central dogma of gene expression.
Transcription
- Main Article: Transcription
During transcription, the gene (double-stranded DNA) is copied or transcribed into a single-stranded messenger RNA molecule (mRNA). In eukaryotes, this process occurs in the nucleus.

Translation: 3) On ribosomes, transfer RNA (tRNA) helps convert mRNA into protein. 4) Amino acids link up to make a protein.
Transcription is very similar to DNA replication although different proteins are involved. The most important enzyme is RNA polymerase, an enzyme that influences the synthesis of RNA from a DNA template. For transcription to be initiated, RNA polymerase must be able to recognize the beginning sequence of a gene so that it knows where to start synthesizing an mRNA. It is directed to this initiation site by the ability of one of its subunits to recognize a specific DNA sequence found at the beginning of a gene, called the promoter sequence. The promoter sequence is a unidirectional sequence found on one strand of the DNA that instructs the RNA polymerase in both where to start synthesis and in which direction synthesis should continue. The RNA polymerase then unwinds the double helix at that point and begins synthesis of a RNA strand complementary to one of the strands of DNA. This strand is called the antisense or template strand, whereas the other strand is referred to as the sense or coding strand. Synthesis can then proceed in a unidirectional manner.[1]
Although much is known about transcript processing, the signals and events that instruct RNA polymerase to stop transcribing and drop off the DNA template remain unclear. Experiments over the years have indicated that processed eukaryotic messages contain a poly(A) addition signal (AAUAAA) at their 3' end, followed by a string of adenines. This poly(A) addition, also called the poly(A) site, contributes not only to the addition of the poly(A) tail but also to transcription termination and the release of RNA polymerase from the DNA template. Yet, transcription does not stop here. Rather, it continues for another 200 to 2000 bases beyond this site before it is aborted. It is either before or during this termination process that the nascent transcript is cleaved, or cut, at the poly(A) site, leading to the creation of two RNA molecules. The upstream portion of the newly formed, or nascent, RNA then undergoes further modifications, called post-transcriptional modification, and becomes mRNA. The downstream RNA becomes unstable and is rapidly degraded.
Although the importance of the poly(A) addition signal has been established, the contribution of sequences further downstream remains uncertain. A recent study suggests that a defined region, called the termination region, is required for proper transcription termination. This study also illustrated that transcription termination takes place in two distinct steps. In the first step, the nascent RNA is cleaved at specific subsections of the termination region, possibly leading to its release from RNA polymerase. In a subsequent step, RNA polymerase disengages from the DNA. Hence, RNA polymerase continues to transcribe the DNA, at least for a short distance.[1]
Translation
- Main Article: Protein synthesis
The message (mRNA) from the gene that is produced by transcription is then carried to an organelle in the cytoplasm (ribosome) where it is translated into a chain of amino acids (polypeptide). The gene is a code that consists of mini coding units called codons, which are 3 nucleotides in length. These codons specify the amino acid that is to be inserted into the growing peptide chain.
| Nucleotide codons and their corresponding amino acids. Codon - Amino Acid TGC = Cysteine CTG = Leucine AGT = Serine GCA = Alanine |
The cellular machinery responsible for synthesizing proteins is the ribosome. The ribosome consists of structural RNA and about 80 different proteins. In its inactive state, it exists as two subunits: a large subunit and a small subunit. When the small subunit encounters an mRNA, the process of translating an mRNA to a protein begins. In the large subunit, there are two sites for amino acids to bind and thus be close enough to each other to form a bond. The "A site" accepts a new transfer RNA, or tRNA—the adaptor molecule that acts as a translator between mRNA and protein—bearing an amino acid. The "P site" binds the tRNA that becomes attached to the growing chain.[1]
The adaptor molecule that acts as a translator between mRNA and protein is a specific RNA molecule, the tRNA. Each tRNA has a specific acceptor site that binds a particular triplet of nucleotides, called a codon, and an anti-codon site that binds a sequence of three unpaired nucleotides, the anti-codon, which can then bind to the the codon. Each tRNA also has a specific charger protein, called an aminoacyl tRNA synthetase. This protein can only bind to that particular tRNA and attach the correct amino acid to the acceptor site.
The start signal for translation is the codon ATG, which codes for methionine. Not every protein necessarily starts with methionine, however. Oftentimes this first amino acid will be removed in later processing of the protein. A tRNA charged with methionine binds to the translation start signal. The large subunit binds to the mRNA and the small subunit, and so begins elongation, the formation of the polypeptide chain. After the first charged tRNA appears in the A site, the ribosome shifts so that the tRNA is now in the P site. New charged tRNAs, corresponding the codons of the mRNA, enter the A site, and a bond is formed between the two amino acids. The first tRNA is now released, and the ribosome shifts again so that a tRNA carrying two amino acids is now in the P site. A new charged tRNA then binds to the A site. This process of elongation continues until the ribosome reaches what is called a stop codon, a triplet of nucleotides that signals the termination of translation. When the ribosome reaches a stop codon, no aminoacyl tRNA binds to the empty A site. This is the ribosome signal to break apart into its large and small subunits, releasing the new protein and the mRNA. Yet, this isn't always the end of the story. A protein will often undergo further modification, called post-translational modification. For example, it might be cleaved by a protein-cutting enzyme, called a protease, at a specific place or have a few of its amino acids altered.[1]
Regulation
- Main Article: regulation of gene expression
Regulation of the gene expression system precisely controls the amount of a gene product that is produced and can further modify the product after it is made. This exquisite control requires multiple regulatory input points. One very efficient point occurs at transcription, such that an mRNA is produced only when a gene product is needed. Cells also regulate gene expression by post-transcriptional modification; by allowing only a subset of the mRNAs to go on to translation; or by restricting translation of specific mRNAs to only when the product is needed. At other levels, cells regulate gene expression through DNA folding, chemical modification of the nucleotide bases, and intricate "feedback mechanisms" in which some of the gene's own protein product directs the cell to cease further protein production.[2]
Gallery
References
- ↑ 1.0 1.1 1.2 1.3 What is a Cell? by the National Center for Biotechnology Information.
- ↑ What is a Genome by the National Center for Biotechnology Information
| |||||||||||||||||